This note is to remind myself about some of the basic concepts of linear regression.
Bibliography
[1] James, Witten, Hastie, Tibshirani - Introduction to Statistical Learning
Setup¶
import numpy as np
import scipy.stats
import pandas as pd
import matplotlib.pylab as plt
%matplotlib inline
(blue, orange, red, green, purple, brown, pink, yellow, lightred, lightblue,
lightorange, lightgreen, lightpurple) = \
('#377eb8', '#ff7f00', '#e41a1c', '#4daf4a', '#984ea3', '#a65628', '#f781bf',
'#d2d215', '#fb9a99', '#a6cee3', '#fdbf6f', '#b2df8a', '#cab2d6')
Creating some (noisy) toy data¶
def create_data(b0, b1, noise_sd, n=100):
"""Return a dataframe with `n` rows and columns 'X' and 'Y'
The dataframe will contain random data points that follow
``Y = b1 * X + b0 + eps`` where the error `eps` is drawn from
a normal distribution with standard deviation `noise_sd`
"""
x = np.sort(10 * np.random.rand(n))
y = b1 * x + b0
noise = scipy.stats.norm(scale=noise_sd)
y += noise.rvs(len(x))
return pd.DataFrame({'X': x, 'Y': y})
b0_true = 0.3
b1_true = 1.2
noise_sd = 2.0 # σ in the standard distribution
data = create_data(b0_true, b1_true, noise_sd)
def plot_data(data, *lines, show=True, **kwargs):
fig, ax = plt.subplots(**kwargs)
data.plot.scatter('X', 'Y', ax=ax)
for (b0, b1, label, color) in lines:
ax.plot(
np.linspace(0, 10, 5),
b0 + np.linspace(0, 10, 5) * b1,
color=color, label=label)
ax.legend()
if show:
plt.show(fig)
else:
return fig, ax
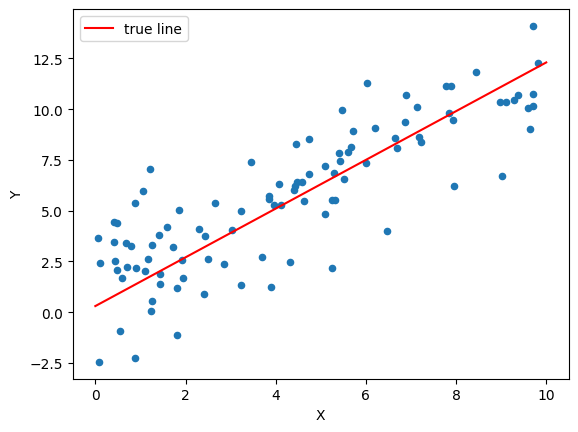
plot_data(data, (b0_true, b1_true, 'true line', 'red'))
Linear Regression with statsmodels¶
import statsmodels.api as sm
mod = sm.OLS(data['Y'], sm.add_constant(data['X']))
res = mod.fit()
res.summary()
res.params
Manual Linear Regression¶
def linear_regression(data):
"""Estimated b0_hat, b1_hat for the given data.
See [1] Eq. (3.4)
"""
xbar = data['X'].mean()
ybar = data['Y'].mean()
delta_x = data['X'] - xbar
delta_y = data['Y'] - ybar
b1_hat = np.dot(delta_x, delta_y) / np.dot(delta_x, delta_x)
b0_hat = ybar - b1_hat * xbar
return b0_hat, b1_hat
b0_hat, b1_hat = linear_regression(data)
b0_hat, b1_hat
b0_true, b1_true
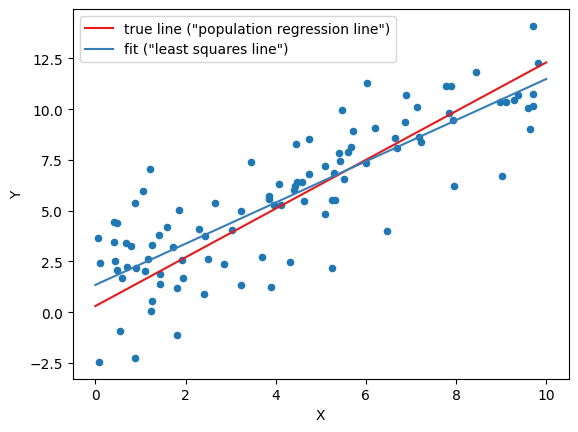
plot_data(
data,
(b0_true, b1_true, 'true line ("population regression line")', red),
(b0_hat, b1_hat, 'fit ("least squares line")', blue)
)
Goodness of Fit¶
Standard Error¶
The Standard Error describes how close an estimated quantity is expected to be to the actual quantity.
It is the standard deviation of the "sampling distribution" of a statistical quantity: If you take a large number of independent samples, each of size $n$, and calculate the statistical quantity for each sample individually, then the standard errror is the standard deviation of the distribution of those results. It is always $\sigma/\sqrt{n}$, where $\sigma$ is the (possibly unknown) standard deviation of the original distribution. Standard errors go to zero as sample sizes go to infinity; standard deviations do not.
def residuals(data, b0, b1):
"""Array of residuals from the perfect linear relation ``Y=b1*X+b0``"""
Y_hat = b1 * data['X'] + b0
return data['Y'] - Y_hat
def RSS(data, b0, b1):
"""Residual Sum of Squares
How much the Y values vary around the predicted Y values
"""
return (residuals(data, b0, b1)**2).sum()
def TSS(data):
"""Total Sum of Squares
How much the Y values vary around the mean Y value
"""
return ((data['Y'] - data['Y'].mean())**2).sum()
def RSE(data, b0, b1):
"""Residual standard error
This is an estimate for the standard deviation σ of the true
distribution (noise_sd). The y-values in the sample deviate from the
from the true regression line by about RSE units on average.
"""
return np.sqrt(RSS(data, b0, b1) / (len(data)-2))
RSS(data, b0_hat, b1_hat)
RSE(data, b0_hat, b1_hat)
noise_sd
def linear_regression_SE(data, sig):
"""Calculate the standard error for the estimate of b0, b1 in a
linear regression for the given data, assuming a standard deviation of
`sig` for the distribution of noise on Y. As `sig` is generally not known,
it should be estimated by the RSE.
Note that the standard error is a property of the sampling only:
The `Y`-values do not enter!
See [1] Eq. (3.8)
"""
n = len(data)
xbar = data['X'].mean()
delta_x = data['X'] - xbar
denom = np.dot(delta_x, delta_x)
se_b0_sq = sig**2 * (1.0 / n + xbar**2/denom)
se_b1_sq = sig**2 / denom
return np.sqrt(se_b0_sq), np.sqrt(se_b1_sq)
linear_regression_SE(data, RSE(data, b0_hat, b1_hat))
The 95% confidence intervals is $\pm$ two standard errors (probability that the true value is in the range of the confidence interval)
Confidence Interval plot¶
See https://stats.stackexchange.com/questions/85560/shape-of-confidence-interval-for-predicted-values-in-linear-regression/85565#85565m for the full explanation and derivation.
def regline_SE(data, sig, xgrid):
"""The standard error of the full regression line, assuming
a standard deviation of `sig` for the distribution of noise on Y. As
`sig` is generally not known, it should be estimated by the RSE.
"""
n = len(data)
return sig * np.sqrt(
(1.0 / n) + (
(xgrid - data['X'].mean())**2 /
((data['X'] - data['X'].mean())**2).sum()))
The 95% confidence band can be calculated as:
def regline_confidence_bands(data, b0_hat, b1_hat, xgrid, confidence=0.95):
"""Return two lines (arrays of y-values), the lower and upper boundary
of the confidence band"""
Y_hat = b0_hat + b1_hat * xgrid # the fitted line, on the given xgrid
n = len(data)
sig = RSE(data, b0_hat, b1_hat)
se = regline_SE(data, sig, xgrid)
t_minus, t_plus = scipy.stats.t(df=n-2).interval(confidence)
return Y_hat + t_minus * se, Y_hat + t_plus * se
fig, ax = plot_data(
data,
(b0_true, b1_true, 'true line ("population regression line")', red),
(b0_hat, b1_hat, 'fit ("least squares line")', blue),
show=False, figsize=(10, 6)
)
xgrid = np.linspace(0, 10, 100)
lm, lp = regline_confidence_bands(data, b0_hat, b1_hat, xgrid)
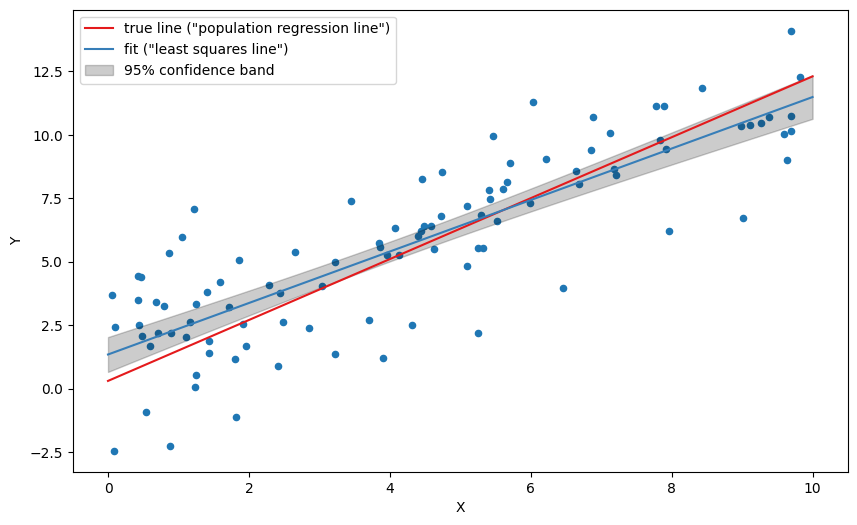
ax.fill_between(
xgrid, lm, lp,
color='black', alpha=0.2, label='95% confidence band')
ax.legend()
plt.show(fig)
The confidence band should be interpreted as follows: There is a 95% probability that a fit based on any given sampling of the same distribution will land in the interval.
Testing the null hypothesis¶
The null-hypthesis is that Y and X have no reltionship, i.e. b1 = 0. The $t$-value is the b1 in units of the standard error:
t = b1_hat / linear_regression_SE(data, RSE(data, b0_hat, b1_hat))[1]
t
The $t$-value follows a student distribution, and we can calculate a p-value (probability that $t$ has the given value, instead of 0)
2 * scipy.stats.t(df=len(data)-2).sf(t) # = 1 - (cdf(t) - cdf(-t))
The $R^2$ statistic¶
def R_sq(data, b0, b1):
"""R^2 in [0, 1] measures how well the variation in Y around the mean
matches the variation in Y around the predicted value.
That is, how much of the variability can be explained by the sampling"""
return 1 - RSS(data, b0, b1) / TSS(data)
R_sq(data, b0_hat, b1_hat)
A value close to 1 means that the linear fit is very good
Normality of the residuals¶
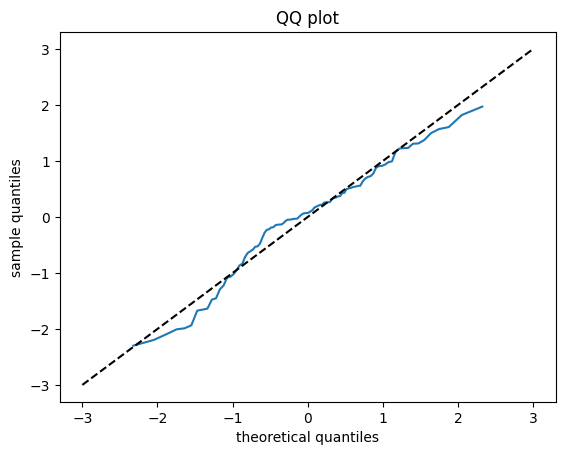
A graphical way to check that the residuals are normal-distributed is to plot the quantiles of the residules against the quantiles of a normal distribution, see https://stats.stackexchange.com/questions/321061/probability-that-residuals-are-normal/321071#321071
fig, ax = plt.subplots()
ax.plot(
scipy.stats.norm().ppf(np.linspace(0, 1, len(data))),
residuals(data, b0_hat, b1_hat).sort_values() / RSE(data, b0_hat, b1_hat))
ax.set_title('QQ plot')
ax.plot(
np.linspace(-3, 3, 10), np.linspace(-3, 3, 10), color='black',
ls='dashed')
ax.set_xlabel('theoretical quantiles')
ax.set_ylabel('sample quantiles')
plt.show(fig)
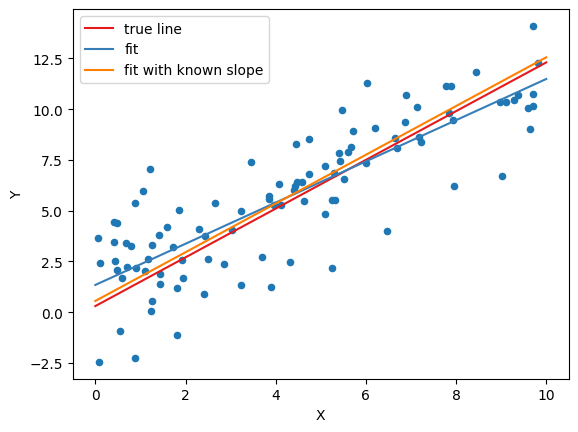
Fit with known slope¶
b0_known_slope = np.mean(data['Y'] - b1_true*data['X'])
b0_known_slope
b0_true
plot_data(
data,
(b0_true, b1_true, 'true line', red),
(b0_hat, b1_hat, 'fit', blue),
(b0_known_slope, b1_true, 'fit with known slope', orange),
show=True,
)